Accuracy¶
Module Interface¶
- class torchmetrics.Accuracy(threshold=0.5, num_classes=None, average='micro', mdmc_average='global', ignore_index=None, top_k=None, multiclass=None, subset_accuracy=False, compute_on_step=None, **kwargs)[source]
Computes Accuracy:
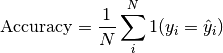
Where
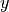 is a tensor of target values, and
is a tensor of target values, and 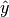 is a
tensor of predictions.
is a
tensor of predictions.For multi-class and multi-dimensional multi-class data with probability or logits predictions, the parameter
top_kgeneralizes this metric to a Top-K accuracy metric: for each sample the top-K highest probability or logit score items are considered to find the correct label.For multi-label and multi-dimensional multi-class inputs, this metric computes the “global” accuracy by default, which counts all labels or sub-samples separately. This can be changed to subset accuracy (which requires all labels or sub-samples in the sample to be correctly predicted) by setting
subset_accuracy=True.Accepts all input types listed in Input types.
- Parameters
num_classes¶ (
Optional[int]) – Number of classes. Necessary for'macro','weighted'andNoneaverage methods.threshold¶ (
float) – Threshold for transforming probability or logit predictions to binary (0,1) predictions, in the case of binary or multi-label inputs. Default value of 0.5 corresponds to input being probabilities.Defines the reduction that is applied. Should be one of the following:
'micro'[default]: Calculate the metric globally, across all samples and classes.'macro': Calculate the metric for each class separately, and average the metrics across classes (with equal weights for each class).'weighted': Calculate the metric for each class separately, and average the metrics across classes, weighting each class by its support (tp + fn).'none'orNone: Calculate the metric for each class separately, and return the metric for every class.'samples': Calculate the metric for each sample, and average the metrics across samples (with equal weights for each sample).
Note
What is considered a sample in the multi-dimensional multi-class case depends on the value of
mdmc_average.Note
If
'none'and a given class doesn’t occur in thepredsortarget, the value for the class will benan.mdmc_average¶ (
Optional[str]) –Defines how averaging is done for multi-dimensional multi-class inputs (on top of the
averageparameter). Should be one of the following:None[default]: Should be left unchanged if your data is not multi-dimensional multi-class.'samplewise': In this case, the statistics are computed separately for each sample on theNaxis, and then averaged over samples. The computation for each sample is done by treating the flattened extra axes...(see Input types) as theNdimension within the sample, and computing the metric for the sample based on that.'global': In this case theNand...dimensions of the inputs (see Input types) are flattened into a newN_Xsample axis, i.e. the inputs are treated as if they were(N_X, C). From here on theaverageparameter applies as usual.
ignore_index¶ (
Optional[int]) – Integer specifying a target class to ignore. If given, this class index does not contribute to the returned score, regardless of reduction method. If an index is ignored, andaverage=Noneor'none', the score for the ignored class will be returned asnan.Number of the highest probability or logit score predictions considered finding the correct label, relevant only for (multi-dimensional) multi-class inputs. The default value (
None) will be interpreted as 1 for these inputs.Should be left at default (
None) for all other types of inputs.multiclass¶ (
Optional[bool]) – Used only in certain special cases, where you want to treat inputs as a different type than what they appear to be. See the parameter’s documentation section for a more detailed explanation and examples.Whether to compute subset accuracy for multi-label and multi-dimensional multi-class inputs (has no effect for other input types).
For multi-label inputs, if the parameter is set to
True, then all labels for each sample must be correctly predicted for the sample to count as correct. If it is set toFalse, then all labels are counted separately - this is equivalent to flattening inputs beforehand (i.e.preds = preds.flatten()and same fortarget).For multi-dimensional multi-class inputs, if the parameter is set to
True, then all sub-sample (on the extra axis) must be correct for the sample to be counted as correct. If it is set toFalse, then all sub-samples are counter separately - this is equivalent, in the case of label predictions, to flattening the inputs beforehand (i.e.preds = preds.flatten()and same fortarget). Note that thetop_kparameter still applies in both cases, if set.
compute_on_step¶ (
Optional[bool]) –Forward only calls
update()and returns None if this is set to False.Deprecated since version v0.8: Argument has no use anymore and will be removed v0.9.
kwargs¶ (
Dict[str,Any]) – Additional keyword arguments, see Advanced metric settings for more info.
- Raises
ValueError – If
top_kis not anintegerlarger than0.ValueError – If
averageis none of"micro","macro","weighted","samples","none",None.ValueError – If two different input modes are provided, eg. using
multi-labelwithmulti-class.ValueError – If
top_kparameter is set formulti-labelinputs.
Example
>>> import torch >>> from torchmetrics import Accuracy >>> target = torch.tensor([0, 1, 2, 3]) >>> preds = torch.tensor([0, 2, 1, 3]) >>> accuracy = Accuracy() >>> accuracy(preds, target) tensor(0.5000)
>>> target = torch.tensor([0, 1, 2]) >>> preds = torch.tensor([[0.1, 0.9, 0], [0.3, 0.1, 0.6], [0.2, 0.5, 0.3]]) >>> accuracy = Accuracy(top_k=2) >>> accuracy(preds, target) tensor(0.6667)
Initializes internal Module state, shared by both nn.Module and ScriptModule.
- compute()[source]
Computes accuracy based on inputs passed in to
updatepreviously.- Return type
Functional Interface¶
- torchmetrics.functional.accuracy(preds, target, average='micro', mdmc_average='global', threshold=0.5, top_k=None, subset_accuracy=False, num_classes=None, multiclass=None, ignore_index=None)[source]
Computes Accuracy
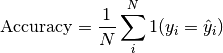
Where
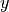 is a tensor of target values, and
is a tensor of target values, and 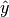 is a
tensor of predictions.
is a
tensor of predictions.For multi-class and multi-dimensional multi-class data with probability or logits predictions, the parameter
top_kgeneralizes this metric to a Top-K accuracy metric: for each sample the top-K highest probability or logits items are considered to find the correct label.For multi-label and multi-dimensional multi-class inputs, this metric computes the “global” accuracy by default, which counts all labels or sub-samples separately. This can be changed to subset accuracy (which requires all labels or sub-samples in the sample to be correctly predicted) by setting
subset_accuracy=True.Accepts all input types listed in Input types.
- Parameters
preds¶ (
Tensor) – Predictions from model (probabilities, logits or labels)Defines the reduction that is applied. Should be one of the following:
'micro'[default]: Calculate the metric globally, across all samples and classes.'macro': Calculate the metric for each class separately, and average the metrics across classes (with equal weights for each class).'weighted': Calculate the metric for each class separately, and average the metrics across classes, weighting each class by its support (tp + fn).'none'orNone: Calculate the metric for each class separately, and return the metric for every class.'samples': Calculate the metric for each sample, and average the metrics across samples (with equal weights for each sample).
Note
What is considered a sample in the multi-dimensional multi-class case depends on the value of
mdmc_average.Note
If
'none'and a given class doesn’t occur in thepredsortarget, the value for the class will benan.mdmc_average¶ (
Optional[str]) –Defines how averaging is done for multi-dimensional multi-class inputs (on top of the
averageparameter). Should be one of the following:None[default]: Should be left unchanged if your data is not multi-dimensional multi-class.'samplewise': In this case, the statistics are computed separately for each sample on theNaxis, and then averaged over samples. The computation for each sample is done by treating the flattened extra axes...(see Input types) as theNdimension within the sample, and computing the metric for the sample based on that.'global': In this case theNand...dimensions of the inputs (see Input types) are flattened into a newN_Xsample axis, i.e. the inputs are treated as if they were(N_X, C). From here on theaverageparameter applies as usual.
num_classes¶ (
Optional[int]) – Number of classes. Necessary for'macro','weighted'andNoneaverage methods.threshold¶ (
float) – Threshold for transforming probability or logit predictions to binary (0,1) predictions, in the case of binary or multi-label inputs. Default value of 0.5 corresponds to input being probabilities.Number of the highest probability or logit score predictions considered finding the correct label, relevant only for (multi-dimensional) multi-class inputs. The default value (
None) will be interpreted as 1 for these inputs.Should be left at default (
None) for all other types of inputs.multiclass¶ (
Optional[bool]) – Used only in certain special cases, where you want to treat inputs as a different type than what they appear to be. See the parameter’s documentation section for a more detailed explanation and examples.ignore_index¶ (
Optional[int]) – Integer specifying a target class to ignore. If given, this class index does not contribute to the returned score, regardless of reduction method. If an index is ignored, andaverage=Noneor'none', the score for the ignored class will be returned asnan.Whether to compute subset accuracy for multi-label and multi-dimensional multi-class inputs (has no effect for other input types).
For multi-label inputs, if the parameter is set to
True, then all labels for each sample must be correctly predicted for the sample to count as correct. If it is set toFalse, then all labels are counted separately - this is equivalent to flattening inputs beforehand (i.e.preds = preds.flatten()and same fortarget).For multi-dimensional multi-class inputs, if the parameter is set to
True, then all sub-sample (on the extra axis) must be correct for the sample to be counted as correct. If it is set toFalse, then all sub-samples are counter separately - this is equivalent, in the case of label predictions, to flattening the inputs beforehand (i.e.preds = preds.flatten()and same fortarget). Note that thetop_kparameter still applies in both cases, if set.
- Raises
ValueError – If
top_kparameter is set formulti-labelinputs.ValueError – If
averageis none of"micro","macro","weighted","samples","none",None.ValueError – If
mdmc_averageis not one ofNone,"samplewise","global".ValueError – If
averageis set butnum_classesis not provided.ValueError – If
num_classesis set andignore_indexis not in the range[0, num_classes).ValueError – If
top_kis not anintegerlarger than0.
Example
>>> import torch >>> from torchmetrics.functional import accuracy >>> target = torch.tensor([0, 1, 2, 3]) >>> preds = torch.tensor([0, 2, 1, 3]) >>> accuracy(preds, target) tensor(0.5000)
>>> target = torch.tensor([0, 1, 2]) >>> preds = torch.tensor([[0.1, 0.9, 0], [0.3, 0.1, 0.6], [0.2, 0.5, 0.3]]) >>> accuracy(preds, target, top_k=2) tensor(0.6667)
- Return type
