Specificity¶
Module Interface¶
- class torchmetrics.Specificity(**kwargs)[source]¶
Compute Specificity.
\[\text{Specificity} = \frac{\text{TN}}{\text{TN} + \text{FP}}\]Where \(\text{TN}\) and \(\text{FP}\) represent the number of true negatives and false positives respectively. The metric is only proper defined when \(\text{TN} + \text{FP} \neq 0\). If this case is encountered for any class/label, the metric for that class/label will be set to 0 and the overall metric may therefore be affected in turn.
This function is a simple wrapper to get the task specific versions of this metric, which is done by setting the
taskargument to either'binary','multiclass'ormultilabel. See the documentation ofBinarySpecificity,MulticlassSpecificityandMultilabelSpecificityfor the specific details of each argument influence and examples.- Legacy Example:
>>> from torch import tensor >>> preds = tensor([2, 0, 2, 1]) >>> target = tensor([1, 1, 2, 0]) >>> specificity = Specificity(task="multiclass", average='macro', num_classes=3) >>> specificity(preds, target) tensor(0.6111) >>> specificity = Specificity(task="multiclass", average='micro', num_classes=3) >>> specificity(preds, target) tensor(0.6250)
BinarySpecificity¶
- class torchmetrics.classification.BinarySpecificity(threshold=0.5, multidim_average='global', ignore_index=None, validate_args=True, **kwargs)[source]¶
Compute Specificity for binary tasks.
\[\text{Specificity} = \frac{\text{TN}}{\text{TN} + \text{FP}}\]Where \(\text{TN}\) and \(\text{FP}\) represent the number of true negatives and false positives respectively. The metric is only proper defined when \(\text{TN} + \text{FP} \neq 0\). If this case is encountered a score of 0 is returned.
As input to
forwardandupdatethe metric accepts the following input:preds(Tensor): An int or float tensor of shape(N, ...). If preds is a floating point tensor with values outside [0,1] range we consider the input to be logits and will auto apply sigmoid per element. Additionally, we convert to int tensor with thresholding using the value inthreshold.target(Tensor): An int tensor of shape(N, ...)
As output to
forwardandcomputethe metric returns the following output:bs(Tensor): Ifmultidim_averageis set toglobal, the metric returns a scalar value. Ifmultidim_averageis set tosamplewise, the metric returns(N,)vector consisting of a scalar value per sample.
If
multidim_averageis set tosamplewisewe expect at least one additional dimension...to be present, which the reduction will then be applied over instead of the sample dimensionN.- Parameters:
threshold¶ (
float) – Threshold for transforming probability to binary {0,1} predictionsmultidim_average¶ (
Literal['global','samplewise']) –Defines how additionally dimensions
...should be handled. Should be one of the following:global: Additional dimensions are flatted along the batch dimensionsamplewise: Statistic will be calculated independently for each sample on theNaxis. The statistics in this case are calculated over the additional dimensions.
ignore_index¶ (
Optional[int]) – Specifies a target value that is ignored and does not contribute to the metric calculationvalidate_args¶ (
bool) – bool indicating if input arguments and tensors should be validated for correctness. Set toFalsefor faster computations.
- Example (preds is int tensor):
>>> from torch import tensor >>> from torchmetrics.classification import BinarySpecificity >>> target = tensor([0, 1, 0, 1, 0, 1]) >>> preds = tensor([0, 0, 1, 1, 0, 1]) >>> metric = BinarySpecificity() >>> metric(preds, target) tensor(0.6667)
- Example (preds is float tensor):
>>> from torchmetrics.classification import BinarySpecificity >>> target = tensor([0, 1, 0, 1, 0, 1]) >>> preds = tensor([0.11, 0.22, 0.84, 0.73, 0.33, 0.92]) >>> metric = BinarySpecificity() >>> metric(preds, target) tensor(0.6667)
- Example (multidim tensors):
>>> from torchmetrics.classification import BinarySpecificity >>> target = tensor([[[0, 1], [1, 0], [0, 1]], [[1, 1], [0, 0], [1, 0]]]) >>> preds = tensor([[[0.59, 0.91], [0.91, 0.99], [0.63, 0.04]], ... [[0.38, 0.04], [0.86, 0.780], [0.45, 0.37]]]) >>> metric = BinarySpecificity(multidim_average='samplewise') >>> metric(preds, target) tensor([0.0000, 0.3333])
- plot(val=None, ax=None)[source]¶
Plot a single or multiple values from the metric.
- Parameters:
val¶ (
Union[Tensor,Sequence[Tensor],None]) – Either a single result from calling metric.forward or metric.compute or a list of these results. If no value is provided, will automatically call metric.compute and plot that result.ax¶ (
Optional[Axes]) – An matplotlib axis object. If provided will add plot to that axis
- Return type:
- Returns:
Figure object and Axes object
- Raises:
ModuleNotFoundError – If matplotlib is not installed
>>> from torch import rand, randint >>> # Example plotting a single value >>> from torchmetrics.classification import BinarySpecificity >>> metric = BinarySpecificity() >>> metric.update(rand(10), randint(2,(10,))) >>> fig_, ax_ = metric.plot()
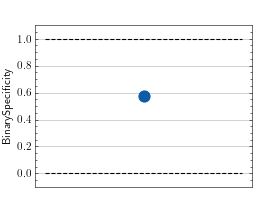
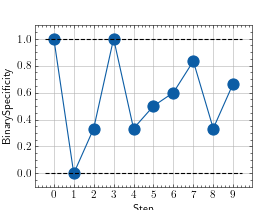
>>> from torch import rand, randint >>> # Example plotting multiple values >>> from torchmetrics.classification import BinarySpecificity >>> metric = BinarySpecificity() >>> values = [ ] >>> for _ in range(10): ... values.append(metric(rand(10), randint(2,(10,)))) >>> fig_, ax_ = metric.plot(values)
MulticlassSpecificity¶
- class torchmetrics.classification.MulticlassSpecificity(num_classes, top_k=1, average='macro', multidim_average='global', ignore_index=None, validate_args=True, **kwargs)[source]¶
Compute Specificity for multiclass tasks.
\[\text{Specificity} = \frac{\text{TN}}{\text{TN} + \text{FP}}\]Where \(\text{TN}\) and \(\text{FP}\) represent the number of true negatives and false positives respectively. The metric is only proper defined when \(\text{TN} + \text{FP} \neq 0\). If this case is encountered for any class, the metric for that class will be set to 0 and the overall metric may therefore be affected in turn.
As input to
forwardandupdatethe metric accepts the following input:preds(Tensor): An int tensor of shape(N, ...)or float tensor of shape(N, C, ..). If preds is a floating point we applytorch.argmaxalong theCdimension to automatically convert probabilities/logits into an int tensor.target(Tensor): An int tensor of shape(N, ...)
As output to
forwardandcomputethe metric returns the following output:mcs(Tensor): The returned shape depends on theaverageandmultidim_averagearguments:If
multidim_averageis set toglobal:If
average='micro'/'macro'/'weighted', the output will be a scalar tensorIf
average=None/'none', the shape will be(C,)
If
multidim_averageis set tosamplewise:If
average='micro'/'macro'/'weighted', the shape will be(N,)If
average=None/'none', the shape will be(N, C)
If
multidim_averageis set tosamplewisewe expect at least one additional dimension...to be present, which the reduction will then be applied over instead of the sample dimensionN.- Parameters:
num_classes¶ (
int) – Integer specifying the number of classesaverage¶ (
Optional[Literal['micro','macro','weighted','none']]) –Defines the reduction that is applied over labels. Should be one of the following:
micro: Sum statistics over all labelsmacro: Calculate statistics for each label and average themweighted: calculates statistics for each label and computes weighted average using their support"none"orNone: calculates statistic for each label and applies no reduction
top_k¶ (
int) – Number of highest probability or logit score predictions considered to find the correct label. Only works whenpredscontain probabilities/logits.multidim_average¶ (
Literal['global','samplewise']) –Defines how additionally dimensions
...should be handled. Should be one of the following:global: Additional dimensions are flatted along the batch dimensionsamplewise: Statistic will be calculated independently for each sample on theNaxis. The statistics in this case are calculated over the additional dimensions.
ignore_index¶ (
Optional[int]) – Specifies a target value that is ignored and does not contribute to the metric calculationvalidate_args¶ (
bool) – bool indicating if input arguments and tensors should be validated for correctness. Set toFalsefor faster computations.
- Example (preds is int tensor):
>>> from torch import tensor >>> from torchmetrics.classification import MulticlassSpecificity >>> target = tensor([2, 1, 0, 0]) >>> preds = tensor([2, 1, 0, 1]) >>> metric = MulticlassSpecificity(num_classes=3) >>> metric(preds, target) tensor(0.8889) >>> mcs = MulticlassSpecificity(num_classes=3, average=None) >>> mcs(preds, target) tensor([1.0000, 0.6667, 1.0000])
- Example (preds is float tensor):
>>> from torchmetrics.classification import MulticlassSpecificity >>> target = tensor([2, 1, 0, 0]) >>> preds = tensor([[0.16, 0.26, 0.58], ... [0.22, 0.61, 0.17], ... [0.71, 0.09, 0.20], ... [0.05, 0.82, 0.13]]) >>> metric = MulticlassSpecificity(num_classes=3) >>> metric(preds, target) tensor(0.8889) >>> mcs = MulticlassSpecificity(num_classes=3, average=None) >>> mcs(preds, target) tensor([1.0000, 0.6667, 1.0000])
- Example (multidim tensors):
>>> from torchmetrics.classification import MulticlassSpecificity >>> target = tensor([[[0, 1], [2, 1], [0, 2]], [[1, 1], [2, 0], [1, 2]]]) >>> preds = tensor([[[0, 2], [2, 0], [0, 1]], [[2, 2], [2, 1], [1, 0]]]) >>> metric = MulticlassSpecificity(num_classes=3, multidim_average='samplewise') >>> metric(preds, target) tensor([0.7500, 0.6556]) >>> mcs = MulticlassSpecificity(num_classes=3, multidim_average='samplewise', average=None) >>> mcs(preds, target) tensor([[0.7500, 0.7500, 0.7500], [0.8000, 0.6667, 0.5000]])
- plot(val=None, ax=None)[source]¶
Plot a single or multiple values from the metric.
- Parameters:
val¶ (
Union[Tensor,Sequence[Tensor],None]) – Either a single result from calling metric.forward or metric.compute or a list of these results. If no value is provided, will automatically call metric.compute and plot that result.ax¶ (
Optional[Axes]) – An matplotlib axis object. If provided will add plot to that axis
- Return type:
- Returns:
Figure object and Axes object
- Raises:
ModuleNotFoundError – If matplotlib is not installed
>>> from torch import randint >>> # Example plotting a single value per class >>> from torchmetrics.classification import MulticlassSpecificity >>> metric = MulticlassSpecificity(num_classes=3, average=None) >>> metric.update(randint(3, (20,)), randint(3, (20,))) >>> fig_, ax_ = metric.plot()
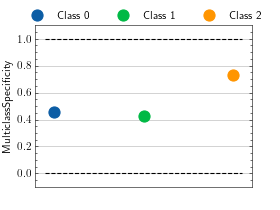
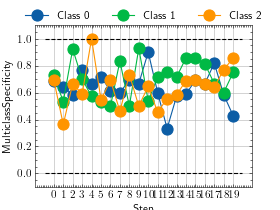
>>> from torch import randint >>> # Example plotting a multiple values per class >>> from torchmetrics.classification import MulticlassSpecificity >>> metric = MulticlassSpecificity(num_classes=3, average=None) >>> values = [] >>> for _ in range(20): ... values.append(metric(randint(3, (20,)), randint(3, (20,)))) >>> fig_, ax_ = metric.plot(values)
MultilabelSpecificity¶
- class torchmetrics.classification.MultilabelSpecificity(num_labels, threshold=0.5, average='macro', multidim_average='global', ignore_index=None, validate_args=True, **kwargs)[source]¶
Compute Specificity for multilabel tasks.
\[\text{Specificity} = \frac{\text{TN}}{\text{TN} + \text{FP}}\]Where \(\text{TN}\) and \(\text{FP}\) represent the number of true negatives and false positives respectively. The metric is only proper defined when \(\text{TN} + \text{FP} \neq 0\). If this case is encountered for any label, the metric for that label will be set to 0 and the overall metric may therefore be affected in turn.
As input to
forwardandupdatethe metric accepts the following input:preds(Tensor): An int or float tensor of shape(N, C, ...). If preds is a floating point tensor with values outside [0,1] range we consider the input to be logits and will auto apply sigmoid per element. Additionally, we convert to int tensor with thresholding using the value inthreshold.target(Tensor): An int tensor of shape(N, C, ...)
As output to
forwardandcomputethe metric returns the following output:mls(Tensor): The returned shape depends on theaverageandmultidim_averagearguments:If
multidim_averageis set toglobalIf
average='micro'/'macro'/'weighted', the output will be a scalar tensorIf
average=None/'none', the shape will be(C,)
If
multidim_averageis set tosamplewiseIf
average='micro'/'macro'/'weighted', the shape will be(N,)If
average=None/'none', the shape will be(N, C)
If
multidim_averageis set tosamplewisewe expect at least one additional dimension...to be present, which the reduction will then be applied over instead of the sample dimensionN.- Parameters:
threshold¶ (
float) – Threshold for transforming probability to binary (0,1) predictionsaverage¶ (
Optional[Literal['micro','macro','weighted','none']]) –Defines the reduction that is applied over labels. Should be one of the following:
micro: Sum statistics over all labelsmacro: Calculate statistics for each label and average themweighted: calculates statistics for each label and computes weighted average using their support"none"orNone: calculates statistic for each label and applies no reduction
multidim_average¶ (
Literal['global','samplewise']) –Defines how additionally dimensions
...should be handled. Should be one of the following:global: Additional dimensions are flatted along the batch dimensionsamplewise: Statistic will be calculated independently for each sample on theNaxis. The statistics in this case are calculated over the additional dimensions.
ignore_index¶ (
Optional[int]) – Specifies a target value that is ignored and does not contribute to the metric calculationvalidate_args¶ (
bool) – bool indicating if input arguments and tensors should be validated for correctness. Set toFalsefor faster computations.
- Example (preds is int tensor):
>>> from torch import tensor >>> from torchmetrics.classification import MultilabelSpecificity >>> target = tensor([[0, 1, 0], [1, 0, 1]]) >>> preds = tensor([[0, 0, 1], [1, 0, 1]]) >>> metric = MultilabelSpecificity(num_labels=3) >>> metric(preds, target) tensor(0.6667) >>> mls = MultilabelSpecificity(num_labels=3, average=None) >>> mls(preds, target) tensor([1., 1., 0.])
- Example (preds is float tensor):
>>> from torchmetrics.classification import MultilabelSpecificity >>> target = tensor([[0, 1, 0], [1, 0, 1]]) >>> preds = tensor([[0.11, 0.22, 0.84], [0.73, 0.33, 0.92]]) >>> metric = MultilabelSpecificity(num_labels=3) >>> metric(preds, target) tensor(0.6667) >>> mls = MultilabelSpecificity(num_labels=3, average=None) >>> mls(preds, target) tensor([1., 1., 0.])
- Example (multidim tensors):
>>> from torchmetrics.classification import MultilabelSpecificity >>> target = tensor([[[0, 1], [1, 0], [0, 1]], [[1, 1], [0, 0], [1, 0]]]) >>> preds = tensor([[[0.59, 0.91], [0.91, 0.99], [0.63, 0.04]], ... [[0.38, 0.04], [0.86, 0.780], [0.45, 0.37]]]) >>> metric = MultilabelSpecificity(num_labels=3, multidim_average='samplewise') >>> metric(preds, target) tensor([0.0000, 0.3333]) >>> mls = MultilabelSpecificity(num_labels=3, multidim_average='samplewise', average=None) >>> mls(preds, target) tensor([[0., 0., 0.], [0., 0., 1.]])
- plot(val=None, ax=None)[source]¶
Plot a single or multiple values from the metric.
- Parameters:
val¶ (
Union[Tensor,Sequence[Tensor],None]) – Either a single result from calling metric.forward or metric.compute or a list of these results. If no value is provided, will automatically call metric.compute and plot that result.ax¶ (
Optional[Axes]) – An matplotlib axis object. If provided will add plot to that axis
- Return type:
- Returns:
Figure object and Axes object
- Raises:
ModuleNotFoundError – If matplotlib is not installed
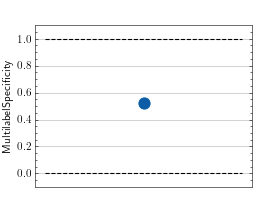
>>> from torch import rand, randint >>> # Example plotting a single value >>> from torchmetrics.classification import MultilabelSpecificity >>> metric = MultilabelSpecificity(num_labels=3) >>> metric.update(randint(2, (20, 3)), randint(2, (20, 3))) >>> fig_, ax_ = metric.plot()
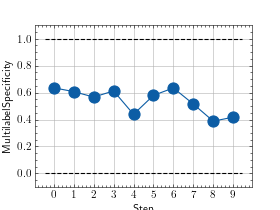
>>> from torch import rand, randint >>> # Example plotting multiple values >>> from torchmetrics.classification import MultilabelSpecificity >>> metric = MultilabelSpecificity(num_labels=3) >>> values = [ ] >>> for _ in range(10): ... values.append(metric(randint(2, (20, 3)), randint(2, (20, 3)))) >>> fig_, ax_ = metric.plot(values)
Functional Interface¶
- torchmetrics.functional.specificity(preds, target, task, threshold=0.5, num_classes=None, num_labels=None, average='micro', multidim_average='global', top_k=1, ignore_index=None, validate_args=True)[source]¶
Compute Specificity. :rtype:
Tensor\[\text{Specificity} = \frac{\text{TN}}{\text{TN} + \text{FP}}\]Where \(\text{TN}\) and \(\text{FP}\) represent the number of true negatives and false positives respecitively.
This function is a simple wrapper to get the task specific versions of this metric, which is done by setting the
taskargument to either'binary','multiclass'ormultilabel. See the documentation ofbinary_specificity(),multiclass_specificity()andmultilabel_specificity()for the specific details of each argument influence and examples.- LegacyExample:
>>> from torch import tensor >>> preds = tensor([2, 0, 2, 1]) >>> target = tensor([1, 1, 2, 0]) >>> specificity(preds, target, task="multiclass", average='macro', num_classes=3) tensor(0.6111) >>> specificity(preds, target, task="multiclass", average='micro', num_classes=3) tensor(0.6250)
binary_specificity¶
- torchmetrics.functional.classification.binary_specificity(preds, target, threshold=0.5, multidim_average='global', ignore_index=None, validate_args=True)[source]¶
Compute Specificity for binary tasks.
\[\text{Specificity} = \frac{\text{TN}}{\text{TN} + \text{FP}}\]Where \(\text{TN}\) and \(\text{FP}\) represent the number of true negatives and false positives respecitively.
Accepts the following input tensors:
preds(int or float tensor):(N, ...). If preds is a floating point tensor with values outside [0,1] range we consider the input to be logits and will auto apply sigmoid per element. Additionally, we convert to int tensor with thresholding using the value inthreshold.target(int tensor):(N, ...)
- Parameters:
threshold¶ (
float) – Threshold for transforming probability to binary {0,1} predictionsmultidim_average¶ (
Literal['global','samplewise']) –Defines how additionally dimensions
...should be handled. Should be one of the following:global: Additional dimensions are flatted along the batch dimensionsamplewise: Statistic will be calculated independently for each sample on theNaxis. The statistics in this case are calculated over the additional dimensions.
ignore_index¶ (
Optional[int]) – Specifies a target value that is ignored and does not contribute to the metric calculationvalidate_args¶ (
bool) – bool indicating if input arguments and tensors should be validated for correctness. Set toFalsefor faster computations.
- Return type:
- Returns:
If
multidim_averageis set toglobal, the metric returns a scalar value. Ifmultidim_averageis set tosamplewise, the metric returns(N,)vector consisting of a scalar value per sample.
- Example (preds is int tensor):
>>> from torch import tensor >>> from torchmetrics.functional.classification import binary_specificity >>> target = tensor([0, 1, 0, 1, 0, 1]) >>> preds = tensor([0, 0, 1, 1, 0, 1]) >>> binary_specificity(preds, target) tensor(0.6667)
- Example (preds is float tensor):
>>> from torchmetrics.functional.classification import binary_specificity >>> target = tensor([0, 1, 0, 1, 0, 1]) >>> preds = tensor([0.11, 0.22, 0.84, 0.73, 0.33, 0.92]) >>> binary_specificity(preds, target) tensor(0.6667)
- Example (multidim tensors):
>>> from torchmetrics.functional.classification import binary_specificity >>> target = tensor([[[0, 1], [1, 0], [0, 1]], [[1, 1], [0, 0], [1, 0]]]) >>> preds = tensor([[[0.59, 0.91], [0.91, 0.99], [0.63, 0.04]], ... [[0.38, 0.04], [0.86, 0.780], [0.45, 0.37]]]) >>> binary_specificity(preds, target, multidim_average='samplewise') tensor([0.0000, 0.3333])
multiclass_specificity¶
- torchmetrics.functional.classification.multiclass_specificity(preds, target, num_classes, average='macro', top_k=1, multidim_average='global', ignore_index=None, validate_args=True)[source]¶
Compute Specificity for multiclass tasks.
\[\text{Specificity} = \frac{\text{TN}}{\text{TN} + \text{FP}}\]Where \(\text{TN}\) and \(\text{FP}\) represent the number of true negatives and false positives respecitively.
Accepts the following input tensors:
preds:(N, ...)(int tensor) or(N, C, ..)(float tensor). If preds is a floating point we applytorch.argmaxalong theCdimension to automatically convert probabilities/logits into an int tensor.target(int tensor):(N, ...)
- Parameters:
num_classes¶ (
int) – Integer specifying the number of classesaverage¶ (
Optional[Literal['micro','macro','weighted','none']]) –Defines the reduction that is applied over labels. Should be one of the following:
micro: Sum statistics over all labelsmacro: Calculate statistics for each label and average themweighted: calculates statistics for each label and computes weighted average using their support"none"orNone: calculates statistic for each label and applies no reduction
top_k¶ (
int) – Number of highest probability or logit score predictions considered to find the correct label. Only works whenpredscontain probabilities/logits.multidim_average¶ (
Literal['global','samplewise']) –Defines how additionally dimensions
...should be handled. Should be one of the following:global: Additional dimensions are flatted along the batch dimensionsamplewise: Statistic will be calculated independently for each sample on theNaxis. The statistics in this case are calculated over the additional dimensions.
ignore_index¶ (
Optional[int]) – Specifies a target value that is ignored and does not contribute to the metric calculationvalidate_args¶ (
bool) – bool indicating if input arguments and tensors should be validated for correctness. Set toFalsefor faster computations.
- Returns:
If
multidim_averageis set toglobal:If
average='micro'/'macro'/'weighted', the output will be a scalar tensorIf
average=None/'none', the shape will be(C,)
If
multidim_averageis set tosamplewise:If
average='micro'/'macro'/'weighted', the shape will be(N,)If
average=None/'none', the shape will be(N, C)
- Return type:
The returned shape depends on the
averageandmultidim_averagearguments
- Example (preds is int tensor):
>>> from torch import tensor >>> from torchmetrics.functional.classification import multiclass_specificity >>> target = tensor([2, 1, 0, 0]) >>> preds = tensor([2, 1, 0, 1]) >>> multiclass_specificity(preds, target, num_classes=3) tensor(0.8889) >>> multiclass_specificity(preds, target, num_classes=3, average=None) tensor([1.0000, 0.6667, 1.0000])
- Example (preds is float tensor):
>>> from torchmetrics.functional.classification import multiclass_specificity >>> target = tensor([2, 1, 0, 0]) >>> preds = tensor([[0.16, 0.26, 0.58], ... [0.22, 0.61, 0.17], ... [0.71, 0.09, 0.20], ... [0.05, 0.82, 0.13]]) >>> multiclass_specificity(preds, target, num_classes=3) tensor(0.8889) >>> multiclass_specificity(preds, target, num_classes=3, average=None) tensor([1.0000, 0.6667, 1.0000])
- Example (multidim tensors):
>>> from torchmetrics.functional.classification import multiclass_specificity >>> target = tensor([[[0, 1], [2, 1], [0, 2]], [[1, 1], [2, 0], [1, 2]]]) >>> preds = tensor([[[0, 2], [2, 0], [0, 1]], [[2, 2], [2, 1], [1, 0]]]) >>> multiclass_specificity(preds, target, num_classes=3, multidim_average='samplewise') tensor([0.7500, 0.6556]) >>> multiclass_specificity(preds, target, num_classes=3, multidim_average='samplewise', average=None) tensor([[0.7500, 0.7500, 0.7500], [0.8000, 0.6667, 0.5000]])
multilabel_specificity¶
- torchmetrics.functional.classification.multilabel_specificity(preds, target, num_labels, threshold=0.5, average='macro', multidim_average='global', ignore_index=None, validate_args=True)[source]¶
Compute Specificity for multilabel tasks.
\[\text{Specificity} = \frac{\text{TN}}{\text{TN} + \text{FP}}\]Where \(\text{TN}\) and \(\text{FP}\) represent the number of true negatives and false positives respecitively.
Accepts the following input tensors:
preds(int or float tensor):(N, C, ...). If preds is a floating point tensor with values outside [0,1] range we consider the input to be logits and will auto apply sigmoid per element. Additionally, we convert to int tensor with thresholding using the value inthreshold.target(int tensor):(N, C, ...)
- Parameters:
threshold¶ (
float) – Threshold for transforming probability to binary (0,1) predictionsaverage¶ (
Optional[Literal['micro','macro','weighted','none']]) –Defines the reduction that is applied over labels. Should be one of the following:
micro: Sum statistics over all labelsmacro: Calculate statistics for each label and average themweighted: calculates statistics for each label and computes weighted average using their support"none"orNone: calculates statistic for each label and applies no reduction
multidim_average¶ (
Literal['global','samplewise']) –Defines how additionally dimensions
...should be handled. Should be one of the following:global: Additional dimensions are flatted along the batch dimensionsamplewise: Statistic will be calculated independently for each sample on theNaxis. The statistics in this case are calculated over the additional dimensions.
ignore_index¶ (
Optional[int]) – Specifies a target value that is ignored and does not contribute to the metric calculationvalidate_args¶ (
bool) – bool indicating if input arguments and tensors should be validated for correctness. Set toFalsefor faster computations.
- Returns:
If
multidim_averageis set toglobal:If
average='micro'/'macro'/'weighted', the output will be a scalar tensorIf
average=None/'none', the shape will be(C,)
If
multidim_averageis set tosamplewise:If
average='micro'/'macro'/'weighted', the shape will be(N,)If
average=None/'none', the shape will be(N, C)
- Return type:
The returned shape depends on the
averageandmultidim_averagearguments
- Example (preds is int tensor):
>>> from torch import tensor >>> from torchmetrics.functional.classification import multilabel_specificity >>> target = tensor([[0, 1, 0], [1, 0, 1]]) >>> preds = tensor([[0, 0, 1], [1, 0, 1]]) >>> multilabel_specificity(preds, target, num_labels=3) tensor(0.6667) >>> multilabel_specificity(preds, target, num_labels=3, average=None) tensor([1., 1., 0.])
- Example (preds is float tensor):
>>> from torchmetrics.functional.classification import multilabel_specificity >>> target = tensor([[0, 1, 0], [1, 0, 1]]) >>> preds = tensor([[0.11, 0.22, 0.84], [0.73, 0.33, 0.92]]) >>> multilabel_specificity(preds, target, num_labels=3) tensor(0.6667) >>> multilabel_specificity(preds, target, num_labels=3, average=None) tensor([1., 1., 0.])
- Example (multidim tensors):
>>> from torchmetrics.functional.classification import multilabel_specificity >>> target = tensor([[[0, 1], [1, 0], [0, 1]], [[1, 1], [0, 0], [1, 0]]]) >>> preds = tensor([[[0.59, 0.91], [0.91, 0.99], [0.63, 0.04]], ... [[0.38, 0.04], [0.86, 0.780], [0.45, 0.37]]]) >>> multilabel_specificity(preds, target, num_labels=3, multidim_average='samplewise') tensor([0.0000, 0.3333]) >>> multilabel_specificity(preds, target, num_labels=3, multidim_average='samplewise', average=None) tensor([[0., 0., 0.], [0., 0., 1.]])
